Information about the journal articles that an institution’s researchers have cited in their own articles can provide valuable data on that institution’s publication record, on the journal usage by researchers as reflected in their publication, and on trends and changes.
The one common denominator in nearly all published citation studies appears to be the difficulty of collecting and evaluating a sufficiently large set of citation data. This article uses an approach that differs from the methods the author found when reviewing a large number of articles on citation analysis. In the majority of articles the data was described as manually compiled from electronic sources [1, 2].
The impetus for the method described in this article was the need to determine the citation use for a specific journal, Mechanisms of Development. To achieve this, the first step was to develop a large data set of references cited by authors at my institution. It was then possible to mine this data set for cited references specific to the target journal.
As a result, the set of data not only answered the initial question about the usage of one journal but formed the basis for other citation-based reports.
The example discussed here highlights several common characteristics of ad-hoc data extraction strategies:
- A good understanding of the quality, scope, and structure of the source data is essential.
- Familiarity with basic strategies of parsing data and selectively extracting nested data. In the example described below regular expressions were not needed, however simple array functions for disassembling the data structure of the exported ISI Web of Science information were critical.
- A clear plan to make sure specific data relationships are preserved in the data output.
- Extracted data sets should provide for data elements that would allow for quick trouble-shooting of the output data. For that purpose the ISI number was included in the output file.
- Subsequent data evaluation and presentation is usually performed more efficiently in common spreadsheet or database programs.
The approach presented in this article is not very technically challenging. Nevertheless, a review of the many articles on citation analysis did not yield any information on similar methods to selectively extract citation information from the ISI citation databases from large exported data sets.
Impetus for the Data Evaluation
Several faculty members at the Medical College of Wisconsin (MCW) expressed a critical need for the journal Mechanisms of Development and suggested that the library purchase a subscription. The high price and the fact that there had not been earlier indications of the importance of the journal led the library to explore ways to determine the need to add a subscription as well as alternatives to a subscription.
It was felt—and the faculty library committee agreed—that for this basic science journal the use in locally published articles would be a good indicator of whether or not a subscription should be purchased. The ISI Web of Science provided the source data.
Obtaining and Examining the Source Data
The source data for this project were collected using the following 2 steps:
-
Performing a search in the ISI Web of Science using a search statement that had been tested previously to ensure the correct set of data would be retrieved. The search criteria in the example include affiliated teaching hospitals.
The search criteria use the Address field in the ISI Web of Science. The institution name(s) were verified using the institution name look-up option in this database.
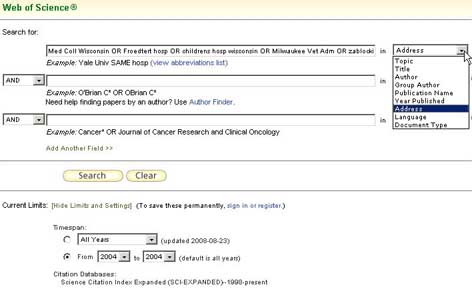
Figure 1: Search Criteria for the Source Data [View full-size image] -
Selecting the appropriate output/export option.
The Perl script used later in this process expects a particular sequence and format of output fields. Below are the settings for the ‘Output Records’ function in the Web of Science that were chosen.
As the number of output records for a project such as the one discussed here will generally exceed the output limit of 500 records, the output procedure will need to be repeated as many times as needed to complete exporting the entire result set.
It is not necessary to redo the search; just the export function will have to be run for each successive set of 500 records. This did not lengthen the time for the data collection significantly.

Figure 2: Output Options [View full-size image]
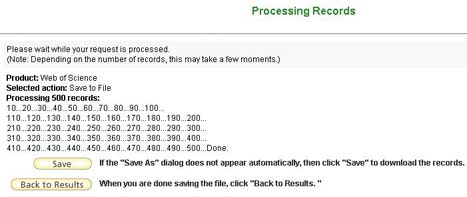
Figure 3: Progress Window from the ISI Web of Science [View full-size image]
The number of records that included the specified institution name(s) varied from year to year between 1,300 to 1,500. From each annual set of exported data about 40,000 cited references were extracted and reported in the script-generated output file.
Examining the Source Data
The example below shows the nested structure of the export format for a full ISI Web of Science citation. The option to include cited references was selected before retrieving the tab-delimited ISI file output format.
Example: (Cited references for this article are in red references to MECH DEVELOP further highlighted in blue.)
J Watt, AJ; Battle, MA; Li, JX; Duncan, SA GATA4 is
essential for formation of the proepicardium and regulates
cardiogenesis PROCEEDINGS OF THE NATIONAL ACADEMY OF
SCIENCES OF THE UNITED STATES OF AMERICA Article
Med Coll Wisconsin, Dept Cell Biol Neurobiol & Anat, Milwaukee,
WI 53202 USA Duncan, SA, Med Coll Wisconsin, Dept Cell Biol
Neurobiol & Anat, Milwaukee, WI 53202 USA duncans@mcw.edu
BRUNEAU BG, 2002, CIRC RES, V90, P509; CHEN THP, 2002, DEV BIOL, V250,
P198; CRIPPS RM, 2002, DEV BIOL, V246, P14; CRISPINO JD, 2001, GENE
DEV, V15, P839; DUNCAN SA, 1997, DEVELOPMENT, V124, P279; DUNWOODIE
SL, 1998, MECH DEVELOP, V72, P27; EISENBERG LM, 1995, CIRC RES, V77,
P1; GARG V, 2003, NATURE, V424, P443; GASSMANN M, 1995, NATURE, V378,
P390; GITTENBERGERDEGROOT AC, 2000, CIRC RES, V87, P969; GREPIN C,
1997, DEVELOPMENT, V124, P2387; JIANG YM, 1996, DEV BIOL, V174, P258;
KISANUKI YY, 2001, DEV BIOL, V230, P230; KUBALAK SW, 1999, HEART DEV,
P209; KUO CT, 1997, GENE DEV, V11, P1048; KWEE L, 1995, DEVELOPMENT,
V121, P489; LEE KF, 1995, NATURE, V378, P394; LIN Q, 1997, SCIENCE,
V276, P1404; LOUGH J, 2000, DEV DYNAM, V217, P327; LYONS I, 1995, GENE
DEV, V9, P1654; MCFADDEN DG, 2000, DEVELOPMENT, V127, P5331; MEYER D,
1995, NATURE, V378, P386; MIKAWA T, 1999, HEART DEV, P19; MJAATVEDT CH,
1999, HEART DEV, P159; MOLKENTIN JD, 1997, GENE DEV, V11, P1061;
MOLKENTIN JD, 2000, J BIOL CHEM, V275, P38949; MOORE AW, 1998, MECH
DEVELOP, V79, P169; NAGY A, 1993, GENE TARGETING PRACT, P147; NARITA N,
1997, DEV BIOL, V189, P270; NARITA N, 1997, DEVELOPMENT, V124, P3755;
NIEDERREITHER K, 2001, DEVELOPMENT, V128, P1019; PARMACEK MS, 1999,
HEART DEV, P291; PARVIZ F, 2003, NAT GENET, V34, P292; PEREZPOMARES JM,
2002, DEV BIOL, V247, P307; ROMEIH M, 2003, DEV DYNAM, V228, P697;
ROSSI JM, 2001, GENE DEV, V15, P1998; SEDMERA D, 1996, BIOESSAYS, V18,
P607; SENGBUSCH JK, 2002, J CELL BIOL, V157, P873; SEPULVEDA JL, 2002,
J BIOL CHEM, V277, P25775; SOUDAIS C, 1995, DEVELOPMENT, V121, P3877;
STENNARD FA, 2003, DEV BIOL, V262, P206; STUCKMANN I, 2003, DEV BIOL,
V255, P334; VIRAGH S, 1973, J ULTRASTRUCT RES, V42, P1; WINNIER G, 1995,
GENE DEV, V9, P2105; YANG JT, 1995, DEVELOPMENT, V121, P549
0027-8424 AUG 24 2004 101 34
12573 12578
ISI:000223596200033
Within the above tab-delimited export of ISI data for an article, the cited references—in red for emphasis—are delimited by semicolons. Each cited reference in turn has elements that are delimited by commas.
Author, Year, Journal, Volume, Page BRUNEAU BG, 2002, CIRC RES, V90, P509; CHEN THP, 2002, DEV BIOL, V250, P198; CRIPPS RM, 2002, DEV BIOL, V246, P14; CRISPINO JD, 2001, GENE DEV, V15, P839; DUNCAN SA, 1997, DEVELOPMENT, V124, P279; DUNWOODIE SL, 1998, MECH DEVELOP, V72, P27; EISENBERG LM, 1995, CIRC RES, V77, P1; (…)
The pattern consists of several layers of nested arrays:
- A nest of Cited Reference Elements—delimited by commas,
- inside a nest of Cited References—delimited by semicolons,
- inside a nest of Data Fields for an Individual Citation—delimited by tabs,
- inside a list of Exported Journal Article Citations from ISI—delimited by End-of Line/Return characters.
Extracting Nested Data Elements with a Perl Script
Virtually any programming or scripting language includes functions to parse input fields in a systematic manner and has methods to handle arrays. Perl was chosen for convenience, and because it is widely used for extracting and transforming data.
The script’s runtime for a typical annual set of data—about 1,500 records—was clocked at than 10 seconds on a PC with a 3.0 GHz Pentium 4.
# PROCESS THE SECTION BELOW THE SCRIPT (BELOW THE __DATA__ TOKEN) # RECORD BY RECORD # file handle for output to a file: $outfile="./isioutrefs2004.txt"; open (OUTPUT, ">$outfile"); while ( <DATA> ) { # remove trailing record separator if found: chomp; # split line by tab delimiter and put the result in an array: @recordline=split (/\t/,$_); $fullrec=$_ ; chomp (@recordline[7]); while ( <DATA> ) { # remove trailing record separator if found: chomp; # split line by tab delimiter and put the result in an array: @recordline=split (/\t/,$_); $fullrec=$_ ; # save the isinumber -the 37th element under a separate variable $isinumber=@recordline[36]; $publishedinjournal=@recordline[4]; chomp (@recordline[7]); foreach (@recordline[14]) { $count++; @citedrefs=split /;/, $_; foreach (@citedrefs) { # do for each cited reference $countrefs++; @onecite=split /,/,$_; # add a line with running number, PubYear, #Title, and ISINumber to the output file print OUTPUT "$countrefs\t$onecite[1]\t$onecite[2]\t$isinumber\n"; # this line just echoes some fields to the screen #as the script runs. # it may be commented out. print "$countrefs\t$onecite[1]\t$onecite[2]\t$isinumber\n"; } } } } # the next two lines just echo some fields to the screen as the script finishes. # they may be commented out. print "Total Refs: $countrefs\n"; print "Total Number of Articles Authored/Co-Authored by MCW: $count"; # the data to be processed begins below the __DATA__ token. __DATA__ PT AU CA TI SO SE LA DT DE ID AB C1 RP EM CR NR TC PU PI PA SN J9 JI PD PY VL IS PN SU SI BP EP AR PG SC GA UT J Watt, AJ; Battle, MA; Li, JX; Duncan, SA GATA4 is essential for formation of the proepicardium and regulates cardiogenesis PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA Article Med Coll Wisconsin, Dept Cell Biol Neurobiol & Anat, Milwaukee, WI 53202 USA Duncan, SA, Med Coll Wisconsin, Dept Cell Biol Neurobiol & Anat, Milwaukee, WI 53202 USA duncans@mcw.edu BRUNEAU BG, 2002, CIRC RES, V90, P509; CHEN THP, 2002, DEV BIOL, V250, P198; CRIPPS RM, 2002, DEV BIOL, V246, P14; CRISPINO JD, 2001, GENE DEV, V15, P839; DUNCAN SA, 1997, DEVELOPMENT, V124, P279; DUNWOODIE SL, 1998, MECH DEVELOP, V72, P27; EISENBERG LM, 1995, CIRC RES, V77, P1; GARG V, 2003, NATURE, V424, P443; GASSMANN M, 1995, NATURE, V378, P390; GITTENBERGERDEGROOT AC, 2000, CIRC RES, V87, P969; GREPIN C, 1997, DEVELOPMENT, V124, P2387; JIANG YM, 1996, DEV BIOL, V174, P258; KISANUKI YY, 2001, DEV BIOL, V230, P230; KUBALAK SW, 1999, HEART DEV, P209; KUO CT, 1997, GENE DEV, V11, P1048; KWEE L, 1995, DEVELOPMENT, V121, P489; LEE KF, 1995, NATURE, V378, P394; LIN Q, 1997, SCIENCE, V276, P1404; LOUGH J, 2000, DEV DYNAM, V217, P327; LYONS I, 1995, GENE DEV, V9, P1654; MCFADDEN DG, 2000, DEVELOPMENT, V127, P5331; MEYER D, 1995, NATURE, V378, P386; MIKAWA T, 1999, HEART DEV, P19; MJAATVEDT CH, 1999, HEART DEV, P159; MOLKENTIN JD, 1997, GENE DEV, V11, P1061; MOLKENTIN JD, 2000, J BIOL CHEM, V275, P38949; MOORE AW, 1998, MECH DEVELOP, V79, P169; NAGY A, 1993, GENE TARGETING PRACT, P147; NARITA N, 1997, DEV BIOL, V189, P270; NARITA N, 1997, DEVELOPMENT, V124, P3755; NIEDERREITHER K, 2001, DEVELOPMENT, V128, P1019; PARMACEK MS, 1999, HEART DEV, P291; PARVIZ F, 2003, NAT GENET, V34, P292; PEREZPOMARES JM, 2002, DEV BIOL, V247, P307; ROMEIH M, 2003, DEV DYNAM, V228, P697; ROSSI JM, 2001, GENE DEV, V15, P1998; SEDMERA D, 1996, BIOESSAYS, V18, P607; SENGBUSCH JK, 2002, J CELL BIOL, V157, P873; SEPULVEDA JL, 2002, J BIOL CHEM, V277, P25775; SOUDAIS C, 1995, DEVELOPMENT, V121, P3877; STENNARD FA, 2003, DEV BIOL, V262, P206; STUCKMANN I, 2003, DEV BIOL, V255, P334; VIRAGH S, 1973, J ULTRASTRUCT RES, V42, P1; WINNIER G, 1995, GENE DEV, V9, P2105; YANG JT, 1995, DEVELOPMENT, V121, P549 0027-8424 AUG 24 2004 101 34 12573 12578 ISI:000223596200033 (…)
Producing Reports from the Data with MS Access
The delimited output file created by running the Perl script can be directly imported into a database such as MS Access. Basic manuals and the ‘Help’ option MS Access provide information about the procedure.
Figure 4 below shows an excerpt from the table in MS Access created after importing the delimited text file that was previously produced by the Perl script.
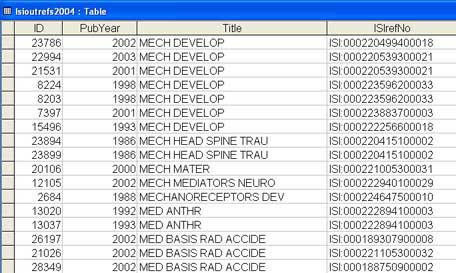
Figure 4: Data Imported into MS Access from the Output of the Perl Script [View full-size image]
To obtain aggregate data, especially counts, the query shown in Figure 5 was composed.
Right-clicking on any column in the query grid shown in Figure 5 will pop up the window that shows ‘Totals’ at the top of the list of choices. After selecting ‘Totals’ the correct option for the ‘Total’ option has to be selected for each column, depending whether the column should be a ‘Group by’ column or contain a specific aggregate function, e.g. Count.
Please note that it is often useful or required to use some columns twice in an aggregate query. The example below uses the ‘Title’ column from the imported table twice: first for displaying the actual journal title in the query output, second for showing the count for the number of times the journal title was found.
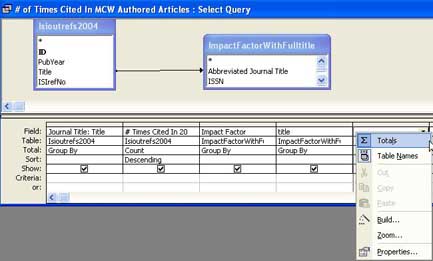
Figure 5: Setting Query Options in MS Access [View full-size image]
A portion of the resulting query output is shown in Figure 6. For the journal, Mechanisms of Development, seven cited references were found in articles authored or co-authored by MCW authors in 2004. Information gathered in the same manner for later years did not show a significant variance in the number from year to year.
During the discussion of this result, the question was raised whether there would be a significant difference in the number of articles that cite a specific journal—perhaps multiple times—versus the overall number of cited references.
Since the ISI number was included in the output file of the Perl script, this number which references a specific article was used to obtain that count.

Figure 6: Query Result for Mechanisms of Development [View full-size image]
For Mechanisms of Development, it was found that the seven cited references were found in five articles. (See Figure 7.)
Figure 7: Query Variation: The number of articles authored/co-authored by MCW Authors in 2004 that cited Mechanisms of Development [View full-size image]
The answer to the question Which journals were cited the most by MCW authors in 2004? was found by sorting the counts. The top of that listing is shown in Figure 8.
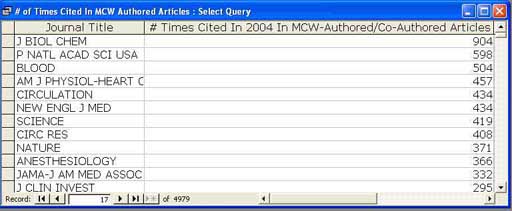
Figure 8: Top of Journal Listing Sorted by Citations in 2004 Articles [View full-size image]
Caveats and Cautions
- The cited reference format in ISI is less regular for references to book chapters, etc. This requires some review of sorted output lists.
- Some journal article references, e.g. those for articles in supplements, differ slightly in the title field. Example: HECK C, 2002, NEUROLOGY S4, V59, S31. This requires caution when counting/grouping references.
- The scope of the ISI databases may not be a suitable match to capture the publishing activities of an institution.
- There are many uses for journal articles that are important but do not result in the publishing of new research.
Outcome / Results
When the data was presented to the faculty library committee, there was agreement that a subscription would not be cost-effective. As an alternative, the library began its pre-paid article service. During the past two years, the request levels for this journal have never exceeded 20% of the subscription price.
The ISI data has become a part of the evaluation of requests for new journals. The trends shown in reports from successive years indicate that dramatic changes from year to year are rare for most journals but noticeable for journals that are making a quick impact in their subject area. It has become evident that the data is valuable as a selection tool for adding new journals as well as for discontinuing subscriptions that are no longer cost-effective, helping keep the selection of journals in alignment with patron needs.
Additional Uses for the Collected Data
Soon after the immediate goal of the data evaluation was met it was realized that the data collected could also yield other journal use metrics. Additional evaluations of the collected data were possible mainly because a comprehensive set of data had been gathered at the start.
Some of the examples of further uses of the data are:
-
Establishing change trends.
A comparison of the journal titles cited by MCW authors with current subscriptions/licenses showed that some heavily-used journals were not among the subscribed journals. When the data for several successive years were included in the comparison, several new journals showed a fast increasing trend line. This additional information—together with faculty requests—was used to adapt the journal subscriptions to the increased relevance of those journals.
Trend information from several successive years is critical for supporting subscription changes.
One key consideration in the decision to add a new journal subscription was the range of years of the cited references. If a currently not subscribed journal is cited increasingly over the recent years and the cited references from the journal are also from recent years the likely cost/benefit ratio of a subscription is more advantageous. This pattern was observed for several—but not all—Nature Reviews journals.
-
Lists of journals that affiliated researcher either publish in or use as a source of cited references.
A sorted listing of journals that researchers primarily rely on for cited references can be produced easily from the collected data. With a small modification of the Perl script, a sorted list of the journals where an institution’s researchers publish can be created.
-
Comparison of counts of cited references vs. articles.
An initial criticism of the output data from the Perl script was that it showed the number of citations from a specific journals used by MCW authors in a particular year but not the number of MCW authored/co-authored articles in which the journal was cited. The concern was that relatively small number of articles could account for a much larger number of cited references to a journal if the articles included many references to articles from the same journal.
Summary
The lack of data extraction and integration methods has impeded the analysis of the scientific communication at a time when the need for a better picture of scientific communication is greater than ever. All participants in the scholarly communication contex—researchers, publishers, academic and other research institutions, etc.—need to better understand measures of quality and effectiveness in scholarly communication.
Analyzing the components of citations/cited references continues to be a foundation for determining the key journals for specific areas of research and for author/institution impact [3].
The method described for extracting data from large collections of citation data shows that a basic but careful evaluation of the source citation data can point to successful data extraction methods that can be quite simple.
Since most citation analyses use sources that also provide their data in either a delimited or—even better—in XML format, there is no reason why citation studies have to be limited to small, manually collected samples.
The author has not yet reviewed other sources for article citation data such as Scopus and Google Scholar. Since the underlying relationships of scholarly communication are the same, the data elements in those sources should be similar.
The method and the basic Perl script presented in this article may be used and modified as needed. Since output formats of the data source—in this instance, the ISI Web of Science—are subject to change, the script may need to be adapted. It is also important to keep in mind that database vendors may restrict specific uses of their data. The license agreement with the database vendor should be consulted.
Notes
1. Glänzel, W. and Moed, H. F. 2002. Journal impact measures in bibliometric research. Scientometrics 53(3): 171-194. COinS
2. Van Raan, A. F. J. 2004. Measuring science. In “Handbook of Quantitative Science and Technology Research. The Use of Publication and Patent Statistics in Studies of S&T Systems” (eds Moed, H. F., Glänzel, W. and Schmoch, U.), Kluwer, Dordrecht, The Netherlands. pp. 19-50. OCLC: 56695465.
3. Meho L. I. Jan. 2007. The rise and rise of citation analysis. Physics World. 20(1): 32-36. Also: arXiv:physics/0701012v1. COinS
About the Author
Alfred Kraemer is Project Manager for academic IT in the Office of Research at the Medical College of Wisconsin in Milwaukee. Until July 2007 he was Assistant Library Director at the same institution.
Author Email: alfred.kraemer@gmail.com


SmartiePants, 2008-09-22
Bravo Alfred, Bravo.
tomkeays, 2010-05-23
ISI Web of Science has changed their output format since the article was published, adding 14 new columns and requiring, therefore, that the Perl script be modified slightly.
In each recordline array:
The full revised script is:
# PROCESS THE SECTION BELOW THE SCRIPT (BELOW THE __DATA__ TOKEN) # RECORD BY RECORD # file handle for output to a file: $outfile="./isioutrefs2004.txt"; open (OUTPUT, ">$outfile"); while ( <DATA> ) { # remove trailing record separator if found: chomp; # split line by tab delimiter and put the result in an array: @recordline=split (/\t/,$_); $fullrec=$_ ; chomp (@recordline[11]); while ( <DATA> ) { # remove trailing record separator if found: chomp; # split line by tab delimiter and put the result in an array: @recordline=split (/\t/,$_); $fullrec=$_ ; # save the isinumber -the 37th element under a separate variable $isinumber=@recordline[50]; $publishedinjournal=@recordline[8]; chomp (@recordline[11]); foreach (@recordline[25]) { $count++; @citedrefs=split /;/, $_; foreach (@citedrefs) { # do for each cited reference $countrefs++; @onecite=split /,/,$_; # add a line with running number, PubYear, #Title, and ISINumber to the output file print OUTPUT "$countrefs\t$onecite[1]\t$onecite[2]\t$isinumber\n"; # this line just echoes some fields to the screen #as the script runs. # it may be commented out. print "$countrefs\t$onecite[1]\t$onecite[2]\t$isinumber\n"; } } } } # the next two lines just echo some fields to the screen as the script finishes. # they may be commented out. print "Total Refs: $countrefs\n"; print "Total Number of Articles Authored/Co-Authored by MCW: $count"; # the data to be processed begins below the __DATA__ token. __DATA__