By Chris Freeland, Martin Kalfatovic, Jay Paige, and Marc Crozier
Introduction
The Biodiversity Heritage Library (BHL)1 is a consortium of ten international natural history museum libraries, botanical garden libraries, and associated technology partners, formed with a mission to digitize the corpus of historic biodiversity literature. The partner libraries include American Museum of Natural History (New York, NY); The Field Museum (Chicago, IL); Harvard University Botany Libraries (Cambridge, MA); Harvard University, Ernst Mayr Library of the Museum of Comparative Zoology (Cambridge, MA); Marine Biological Laboratory / Woods Hole Oceanographic Institution (Woods Hole, MA); Missouri Botanical Garden (St. Louis, MO); Natural History Museum (London, UK); The New York Botanical Garden (New York, NY); Royal Botanic Gardens, Kew (Richmond, UK); and the Smithsonian Institution Libraries (Washington, DC). The University Library of the University of Illinois at Urbana-Champaign (Urbana, IL), a lead member of the Open Content Alliance, is also supporting the work of BHL as a Contributing member.
The collections held by BHL partners total upwards of two million volumes and contain core scientific literature dating from the 15th century. These collections have been curated for more than 200 years to support the research activities of scientists and students describing the living world. Because of shared interest with smaller scientific societies, BHL collections offer many unique titles unavailable even in larger university collections. From a scholarly perspective, these collections are of exceptional value because the domain of systematic biology depends upon references to original publication of species descriptions contained in historic literature.2
However, this wealth of knowledge is readily available to only a small subset of the entire community who need access to the materials, as these collections are primarily held in large libraries in North America and Europe. Scientists must spend considerable time and funds to travel to these libraries for their research. Further, while in non-digital form, this corpus of literature is unavailable to data mining endeavors within taxonomic study, biodiversity conservation, protected area management, disease control, and associated educational outreach.
In addition to works in the public domain, BHL is committed to working with partners in the scholarly publishing community to obtain permission to digitize materials of scientific interest and make it available in an open access environment. This “opt in” model for digitization will allow for the broadest representation of taxonomic literature while obtaining buy-in from the scholarly publishing world.
Digitization and Metadata Creation
In 2005, concurrent with early BHL planning sessions, the Missouri Botanical Garden (MOBOT) received funding from the W.M. Keck Foundation to model a robust software platform to connect biodiversity databases with digital library content, including interfaces, both user-centered and programmatic, that addressed the specific needs of the taxonomic research community.3 The platform, named Botanicus, was demonstrated using materials scanned in-house at the Missouri Botanical Garden and is online at www.botanicus.org.
BHL project planning began in earnest during 2006. The Internet Archive (IA)4 was invited to join BHL as a Technical Partner because of its efficient, high volume Scribe scanning workstations and commitment to open access distribution of public domain materials through its lead participation in the Open Content Alliance.5 In June of 2007 BHL received funding from the John D. and Catherine T. MacArthur Foundation and the Alfred P. Sloan Foundation to scan upwards of 60 million pages of scientific literature held across BHL partner collections. Book digitization began in late summer of 2007 at scanning centers in London, Boston, New York, Washington DC, and at the University of Illinois Urbana-Champaign. Botanicus was adopted as the model for BHL, with slight modifications, and deployed at www.biodiversitylibrary.org.
At the time of scanning, the MARC record for the monograph or serial being scanned is retrieved from the contributing library’s ILS using a YAZ Z39.50 fetch6 and transformed to MARCXML. The MARC and MARCXML records are deposited alongside the raw camera images in a directory for each scanned book on IA clusters. Additional structural and administrative metadata, created at the time of scanning, is also deposited in the book’s directory. Automated processes run against the raw files to produce a series of derivatives, including JPEG2000 images, color PDF, black and white PDF, and OCR text.7
To facilitate the advanced features and functionality modeled for the taxonomic community through development of the BHL portal, the files for a scanned monograph or serial volume are ingested from IA clusters and transformed into the BHL data model. In this way, books and bibliographic metadata from multiple libraries are delivered through a common interface with advanced functionality specifically developed for the communities served by BHL partner libraries.
Metadata Reuse
BHL partners were particularly keen to reuse the vast amounts of specialized descriptive metadata created by professional catalogers contained in the bibliographic records. Partners were also interested in evaluating new visualization techniques for displaying these data in a modern, service-oriented interface. Library of Congress Subject Headings (LCSH) were selected as the first dataset for experimentation as they offer a controlled vocabulary that has been routinely applied within BHL libraries, and which offer an important access point for users looking for materials on a given topic.
Because the assignment of LCSH is regimented, these data provided an interesting counterpoint to the popular application of user-generated tags, also known as a folksonomy.8 Collections of user-generated tags are often displayed in tag clouds, which give visual weight and emphasis to tags representing more content and deemphasize tags representing less content, as demonstrated in Figure 1 from the photo sharing portal Flickr.
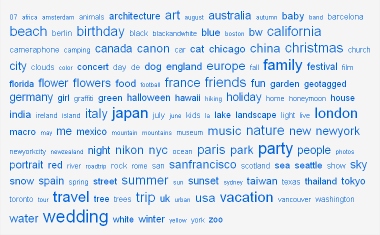 Figure 1: “All time most popular tags” from Flickr, 19 Jan 2008. http://www.flickr.com/photos/tags/. [View full-size image]
Figure 1: “All time most popular tags” from Flickr, 19 Jan 2008. http://www.flickr.com/photos/tags/. [View full-size image]
Another example is the Internet Archive, which displays a tag cloud for its digitized content, including its Children’s Book Collection.9
To build a similar interface from LCSH, the MARCXML for each scanned title is parsed into corresponding fields in the BHL database, currently deployed in SQL Server 2005 on servers at the Missouri Botanical Garden. Working under the guidance of librarians at BHL partner libraries, all subject headings (MARC 6XX fields)10 available in a given record are selected to form the tag cloud.
This activity commits each subdivision of a LCSH string to its own database record associated with the title from which it was assigned. These records are then grouped according to their number of occurrences and displayed using a tag cloud interface.
For a digital library the size of BHL, currently expected to include more than 60 million scanned pages by 2012, techniques are required to selectively view subject tags, as viewing the entire cloud would require several seconds to draw and would include hundreds of results with only a single associated title. Figure 2 illustrates the approach taken with the BHL portal. By default, users are presented with the Top 100 subjects and given links to other quantities, as well as a link to view all tags. This approach allows the tag cloud and surrounding page to draw quickly, while still providing interfaces to those who want to view more details.
 Figure 2: Tag cloud from BHL, 25 Feb 2008. http://www.biodiversitylibrary.org/browse/subject. [View full-size image]
Figure 2: Tag cloud from BHL, 25 Feb 2008. http://www.biodiversitylibrary.org/browse/subject. [View full-size image]
Geocoding LCSH
To then further extend use of LCSH, each term recorded in subfield $z (geographic subdivision) is geocoded using classes and methods within the Google Maps Application Programming Interface (API).11 Geocoding is the process by which structured geospatial information is determined for map-based data, such as coordinates for a street addresses or, in this example, coordinates for a geographic place name.12 The resulting interface is displayed within the BHL Portal at http://www.biodiversitylibrary.org/browse/map and in Figure 3.
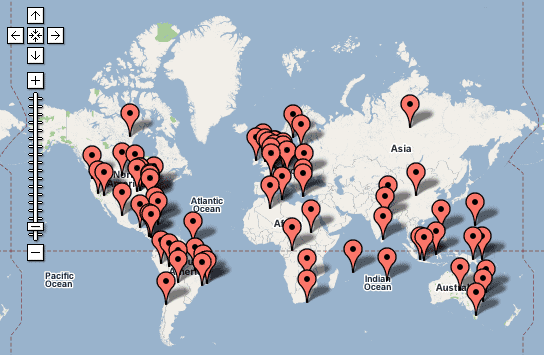 Figure 3: Google Map with geocoded LCSH from the Biodiversity Heritage Library.
Figure 3: Google Map with geocoded LCSH from the Biodiversity Heritage Library.
To begin using the Google Maps API for geocoding, organizations must first sign up for an API key that is submitted along with each API call.13 The Google Maps API geocoder is accessible via several protocols and can return results in a variety of formats, including CSV, XML, Google KML, and JSON.14 In current practice, BHL submits strings for geocoding via HTTP from server code and receives responses in CSV, as these methods are the easiest to form and least bandwidth-intensive. Here is a sample HTTP geocode request for the tag “Brazil” (with parameters on separate lines for clarity):
http://maps.google.com/maps/geo?output=csv &key=[developer api key] &q=Brazil
This request returns the following result:
200,1,-14.235004,-51.925280
“200” is the HTTP status code (which indicates success)
“1” indicates accuracy (in this case, “country” level accuracy)
“-14.235004” is the latitude
“-51.925280” is the longitude
As a comparison, here is the same result when formatted in KML, Google’s XML-based language for geospatial data:
<?xml version="1.0" encoding="UTF-8"?> <kml xmlns="http://earth.google.com/kml/2.0"> <Response> <name>Brazil</name> <Status> <code>200</code> <request>geocode</request> </Status> <Placemark id="p1"> <address>Brazil</address> <AddressDetails Accuracy="1" xmlns="urn:oasis:names:tc:ciq:xsdschema:xAL:2.0"> <Country> <CountryNameCode>BR</CountryNameCode> </Country> </AddressDetails> <Point> <coordinates>-51.925280,-14.235004,0</coordinates> </Point> </Placemark> </Response> </kml>
Regardless of response format, the values returned for latitude and longitude are stored in the BHL database. If geocoding was unsuccessful, those tags are marked as a failure in the database with a timestamp to re-attempt the geocode after 30 days. This way each failed tag is resubmitted at regular intervals, allowing for the possibility that Google may be able to find it in the future should their list of place names or their geocoding algorithms change.
When a user browses to the page at http://www.biodiversitylibrary.org/browse/map, server-side code first looks to see if there are any new tags that need to be geocoded, since new tags are added to BHL as new materials are ingested. After submitting new tags for geocoding, the tags that have previously failed are submitted again if it is 30 days after their last attempt. This additional lookup is negligible and does not affect response times or user experience. Finally, all tags with latitude and longitude are sent to the browser and are drawn on the map. In this way values can be stored and cached for increased performance, while still allowing updates to occur for new tags and tags that have previously failed. Another approach would be to schedule this lookup as a cron job or other scheduled task on the server, but BHL developers favored including the lookup directly in the web application itself in order to minimize the number of dependent applications.
BHL is not alone in visualizing LCSH through tag clouds or Google Maps. “The Maritime History of the Great Lakes” digital collection uses similar techniques to display LCSH in a tag cloud,15 as well as geocoding LCSH and displaying a map for individual records.16
This presentation and functionality has been well-received by our user community, who have described the interface as “a system to quickly and easily find the information you want with minimal distractions.”17
Complications and Limitations
Although successful, this method of geocoding and presenting data gleaned from LCSH is not without problems. Some of the issues are in the data themselves and some have to do with limitations of the Google Maps API and the methods by which it geocodes and displays Placemarks.
The main issue concerning LCSH values is that they are only as complete as each BHL partner’s cataloging practices. BHL partners are not explicitly applying or improving LCSH before scanning, so if there are no values present in the original MARC record there are no data to geocode, leading to visual gaps in the display. For example, a BHL partner might have digitized a volume about Costa Rica, but if that assertion has not been recorded in LCSH there will not be a Placemark on Costa Rica in the map display.
Other limitations in the display relate to how Google Maps geocodes values and how it then displays those Placemarks. By definition a Placemark is a single point on a map, which works well with traditional uses of Google Maps, such as displaying points for street addresses. However, geocoding returns a somewhat different result for less granular place names like “Missouri.” The point associated with “Missouri” is the centroid, or center point, of the polygon defined by the boundaries of the state of Missouri. While this is not a major barrier to viewing results, it does pose some ambiguity when there are Placemarks for both a state and a city within the state, such as “Missouri” and “St. Louis”, as demonstrated in Figure 4.
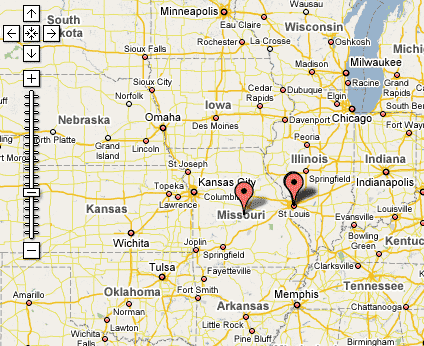 Figure 4: Google Map showing Placemarks for “Missouri” and “St. Louis”.
Figure 4: Google Map showing Placemarks for “Missouri” and “St. Louis”.
Even as place names describe larger geographic areas, such as continents, a single Placemark is plotted on the centroid. Figure 5 shows how the place name “South America” is plotted in southern Bolivia, as that point represents the center of the geographic boundaries of the continent. As demonstrated by these two examples, a single Placemark for a regional place name might produce ambiguous results when plotted alongside more specific place names.
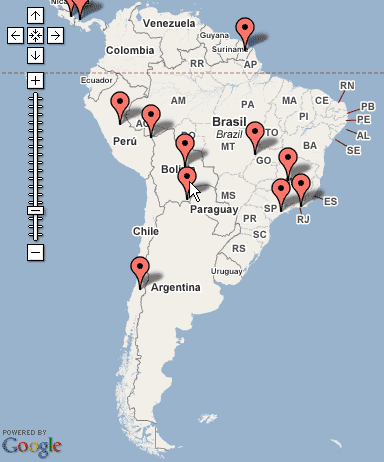 Figure 5: Placemark for “South America” as indicated by pointer.
Figure 5: Placemark for “South America” as indicated by pointer.
A further complication with the single Placemark paradigm relates to the inability to visually represent areas of density within Google Maps. Rather than viewing single Placemarks at each country’s centroid, a more compelling display would be to view a map with shading to represent countries associated with more digitized content, similar in display to a population density map. It is possible to cluster Placemarks such that multiple points are represented by a single Placemark when zoomed out on the map,18 allowing developers to streamline maps with many points in close proximity to one another. However, clustering still doesn’t allow for visual ranking or weighting of results.
Unfortunately the only methods within Google Maps to define polygons (such as state or continental boundaries) and color-code for visual weight are cumbersome to use and difficult to display without considerable delays in performance. To accurately draw the polygon for South America each point along the continent’s coastline would have to be defined, requiring thousands of points and therefore significant delays in delivering the map to users (upwards of 20 seconds or longer each time the map is drawn). An illustration of this point is available by viewing voting results from the 2008 New Hampshire Primaries.19
Future Work
In addition to investigating techniques for displaying polygons and relative density, BHL developers plan to evaluate other mapping services. Alternatives to Google Maps, such as Yahoo! Maps Web Services and Microsoft Virtual Earth, have grown in functionality and community use. These services, as well as other efforts underway within the OpenGIS community, bear further evaluation. Work is also needed to perform a more thorough study of user experience and user behavior to determine efficacy of browsing a digital library through a map-based interface.
Another area of further investigation is to geocode and map MARC values outside of LCSH, such as the MARC Code List for Geographic Areas, commonly represented within MARC field 043.20 The same complications with incomplete data exist within this field, similar to the problem previously stated with LCSH.
Conclusion
Reusing metadata generated through years of cataloging practice is a natural and pragmatic way of leveraging an institution’s investment in describing its resources. Unfortunately, many digitization projects disassociate metadata about the original object from the metadata about its digital surrogate when publishing that surrogate online.
Through reuse of LCSH in forming tag clouds and interactive maps, BHL has demonstrated how easily that original cataloging work can be brought into the web space and partnered with emerging technology to provide advanced user interfaces and functionality. While the methods described in this paper result in access points that are not comprehensive, they do provide new methods of resource discovery and new opportunities for evaluating visualization techniques. It is of utmost importance that libraries continually evaluate how best to reuse legacy data in light of emerging technologies and to determine how best to remix those data into new and innovative interfaces.
Notes
1. http://www.biodiversitylibrary.org
2. Krell, Frank-Thorsten. 2000. “Impact factors aren’t relevant to taxonomy.” Nature. 405: 507-508. doi:10.1038/35014664. COinS
3. http://www.botanicus.org/About.aspx
5. http://www.opencontentalliance.org/
6. http://www.indexdata.com/yaz/
7. http://www.archive.org/download/freshwaterfishes00eige/
8. http://en.wikipedia.org/wiki/Folksonomy
9. http://www.archive.org/browse.php?field=/metadata/subject&collection=iacl&view=cloud
10. http://www.loc.gov/marc/bibliographic/ecbdsubj.html
11. http://code.google.com/apis/maps/documentation/reference.html
12. http://en.wikipedia.org/wiki/Geocoding
13. http://code.google.com/apis/maps/signup.html
14. http://code.google.com/apis/maps/documentation/services.html#Geocoding_Direct
15. http://www.hhpl.on.ca/GreatLakes/GLImages/BrowseBySubject.asp
16. http://www.hhpl.on.ca/GreatLakes/GLImages/details.asp?ID=9
17. http://moultriecreek.us/family/2007/11/03/is-this-any-way-to-run-an-archive/trackback/
18. http://googlemapsapi.110mb.com/clustermarker/index.htm
19. http://maps.google.com/maps/mpl?moduleurl=
http:%2F%2Fwww.google.com%2Fmapfiles%2Fmapplets%2Fnhprimary%2Fnhprimary.xml
&ie=UTF8&om=0&ll=43.975,-71.628&spn=1.585195,3.691406&z=8
20. http://www.loc.gov/marc/geoareas/gacs_code.pdf
About the Authors
Chris Freeland is the Technical Director of the Biodiversity Heritage Library and manager of the Bioinformatics department at the Missouri Botanical Garden.
Martin Kalfatovic is Head of the New Media Office and Preservation Services at Smithsonian Institution Libraries.
Jay Paige is a Senior Programmer in the Missouri Botanical Garden’s Bioinformatics department.
Marc Crozier is a programming consultant in the Missouri Botanical Garden’s Bioinformatics department.


Chris Freeland, 2008-03-26
If you are interested in learning more about the process for ingesting and parsing content from Internet Archive, read the BHL Blog entry on the topic.
Jonathan Rochkind, 2008-05-20
I just discovered this open access source of place name to lat/long mapping, essentially an OA gazeteer.
http://www.geonames.org/
Super nice!
Yes, it has a web service, as well as downloadable data. Seems worth considering for any projects that need to translate from existing controlled place vocabulary without lat/long (like LCSH!) to lat/long.
Thoughts on “Geocoding in the LCSH Biodiversity Library” « Digital Mappings: Readings & Reflections, 2012-05-16
[…] Geocoding in the LCSH Biodiversity Library, the authors detail how they have taken MARC record locations and ingested them through the Google […]
regeniaec, 2012-05-22
A fantastic view of point for google maps. Never miss a thing when you need it most of the time to find a location that your not familiar with.